step_rose() creates a specification of a recipe step that generates
samples of synthetic data by enlarging the feature space of minority and
majority class examples. Using ROSE::ROSE().
Usage
step_rose(
recipe,
...,
role = NA,
trained = FALSE,
column = NULL,
over_ratio = 1,
minority_prop = 0.5,
minority_smoothness = 1,
majority_smoothness = 1,
indicator_column = NULL,
skip = TRUE,
seed = sample.int(10^5, 1),
id = rand_id("rose")
)Arguments
- recipe
A recipe object. The step will be added to the sequence of operations for this recipe.
- ...
One or more selector functions to choose which variable is used to sample the data. See recipes::selections for more details. The selection should result in single factor variable. For the
tidymethod, these are not currently used.- role
Not used by this step since no new variables are created.
- trained
A logical to indicate if the quantities for preprocessing have been estimated.
- column
A character string of the variable name that will be populated (eventually) by the
...selectors.- over_ratio
A numeric value for the ratio of the minority-to-majority frequencies. The default value (1) means that all other levels are sampled up to have the same frequency as the most occurring level. A value of 0.5 would mean that the minority levels will have (at most) (approximately) half as many rows as the majority level. See
vignette("ratio", package = "themis")for more details.- minority_prop
A numeric value between 0 and 1 for the proportion of synthetic observations from the minority class. Defaults to 0.5, which generates an equal split of minority and majority synthetic observations. This parameter controls the class balance within the synthetic data, while
over_ratiocontrols the total size of the synthetic data.- minority_smoothness
A numeric. Shrink factor to be multiplied by the smoothing parameters to estimate the conditional kernel density of the minority class. Defaults to 1.
- majority_smoothness
A numeric. Shrink factor to be multiplied by the smoothing parameters to estimate the conditional kernel density of the majority class. Defaults to 1.
- indicator_column
A single string or
NULL(the default). If a string is given, a logical column with that name is added to the output. Because ROSE generates a fully synthetic dataset, all rows are markedTRUE.- skip
A logical. Should the step be skipped when the recipe is baked by
bake()? While all operations are baked whenprep()is run, some operations may not be able to be conducted on new data (e.g. processing the outcome variable(s)). Care should be taken when usingskip = TRUEas it may affect the computations for subsequent operations.- seed
An integer that will be used as the seed when rose-ing.
- id
A character string that is unique to this step to identify it.
Value
An updated version of recipe with the new step
added to the sequence of existing steps (if any). For the
tidy method, a tibble with columns terms which is
the variable used to sample.
Details
The factor variable used to balance around must only have 2 levels.
The ROSE algorithm works by selecting an observation belonging to class k
and generating new examples in its neighborhood, which is determined by a
smoothing matrix H_k. Smaller values of minority_smoothness and
majority_smoothness shrink the entries of H_k, producing tighter
neighborhoods. This is a cautious choice when there is a concern that
excessively large neighborhoods could blur the boundaries between classes.
All columns in the data are sampled and returned by recipes::juice()
and recipes::bake().
When used in modeling, users should strongly consider using the
option skip = TRUE so that the extra sampling is not
conducted outside of the training set.
Tidying
When you tidy() this step, a tibble is returned with
columns terms and id:
- terms
character, the selectors or variables selected
- id
character, id of this step
Tuning Parameters
This step has 1 tuning parameters:
over_ratio: Over-Sampling Ratio (type: double, default: 1)
References
Lunardon, N., Menardi, G., and Torelli, N. (2014). ROSE: a Package for Binary Imbalanced Learning. R Journal, 6:82–92.
Menardi, G. and Torelli, N. (2014). Training and assessing classification rules with imbalanced data. Data Mining and Knowledge Discovery, 28:92–122.
See also
rose() for direct implementation
Other Steps for over-sampling:
step_adasyn(),
step_bsmote(),
step_smote(),
step_smotenc(),
step_upsample()
Examples
library(recipes)
library(modeldata)
data(hpc_data)
hpc_data0 <- hpc_data |>
mutate(class = factor(class == "VF", labels = c("not VF", "VF"))) |>
select(-protocol, -day)
orig <- count(hpc_data0, class, name = "orig")
orig
#> # A tibble: 2 × 2
#> class orig
#> <fct> <int>
#> 1 not VF 2120
#> 2 VF 2211
up_rec <- recipe(class ~ ., data = hpc_data0) |>
step_rose(class) |>
prep()
training <- up_rec |>
bake(new_data = NULL) |>
count(class, name = "training")
training
#> # A tibble: 2 × 2
#> class training
#> <fct> <int>
#> 1 not VF 2242
#> 2 VF 2180
# Since `skip` defaults to TRUE, baking the step has no effect
baked <- up_rec |>
bake(new_data = hpc_data0) |>
count(class, name = "baked")
baked
#> # A tibble: 2 × 2
#> class baked
#> <fct> <int>
#> 1 not VF 2120
#> 2 VF 2211
orig |>
left_join(training, by = "class") |>
left_join(baked, by = "class")
#> # A tibble: 2 × 4
#> class orig training baked
#> <fct> <int> <int> <int>
#> 1 not VF 2120 2242 2120
#> 2 VF 2211 2180 2211
library(ggplot2)
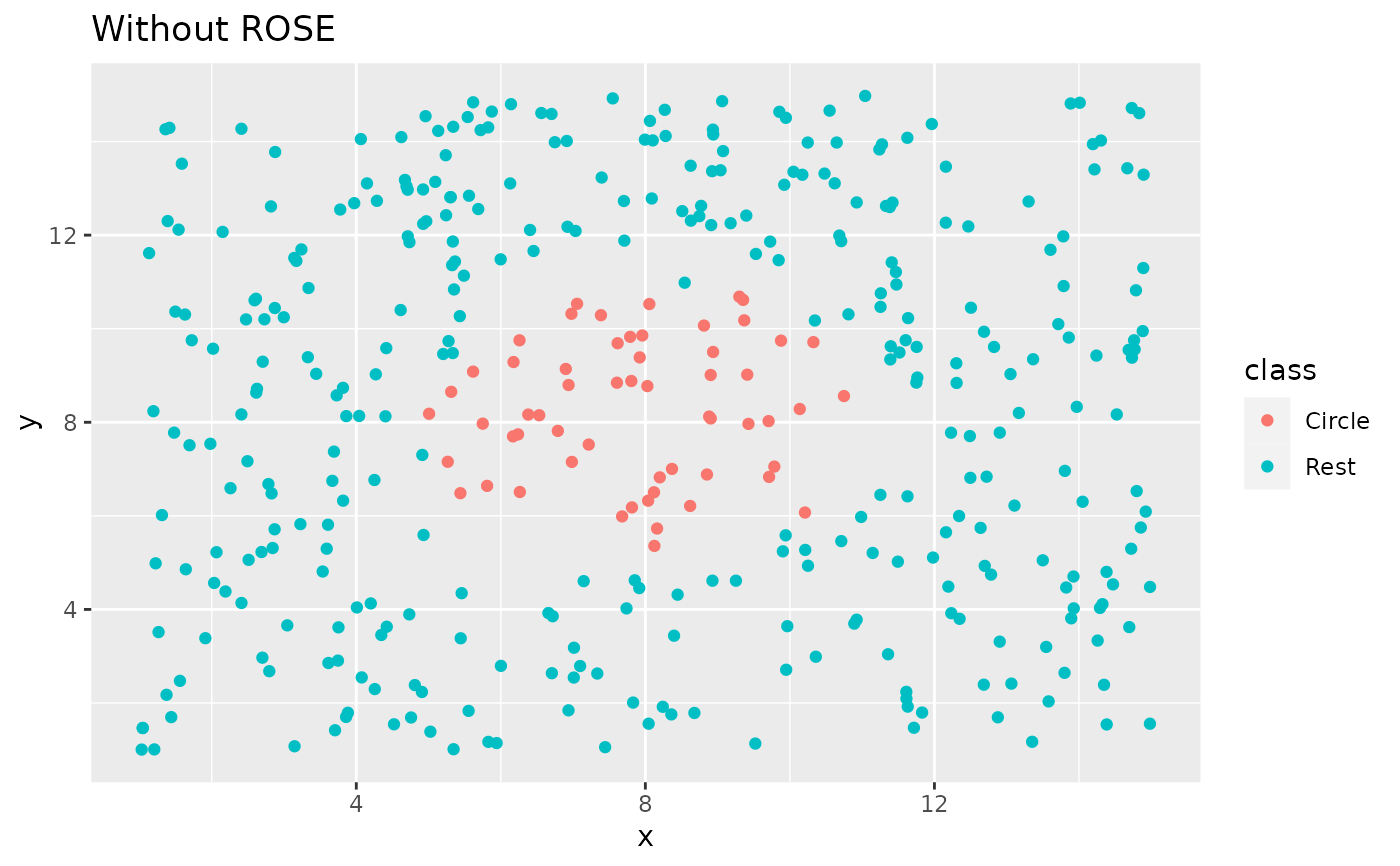
ggplot(circle_example, aes(x, y, color = class)) +
geom_point() +
labs(title = "Without ROSE")
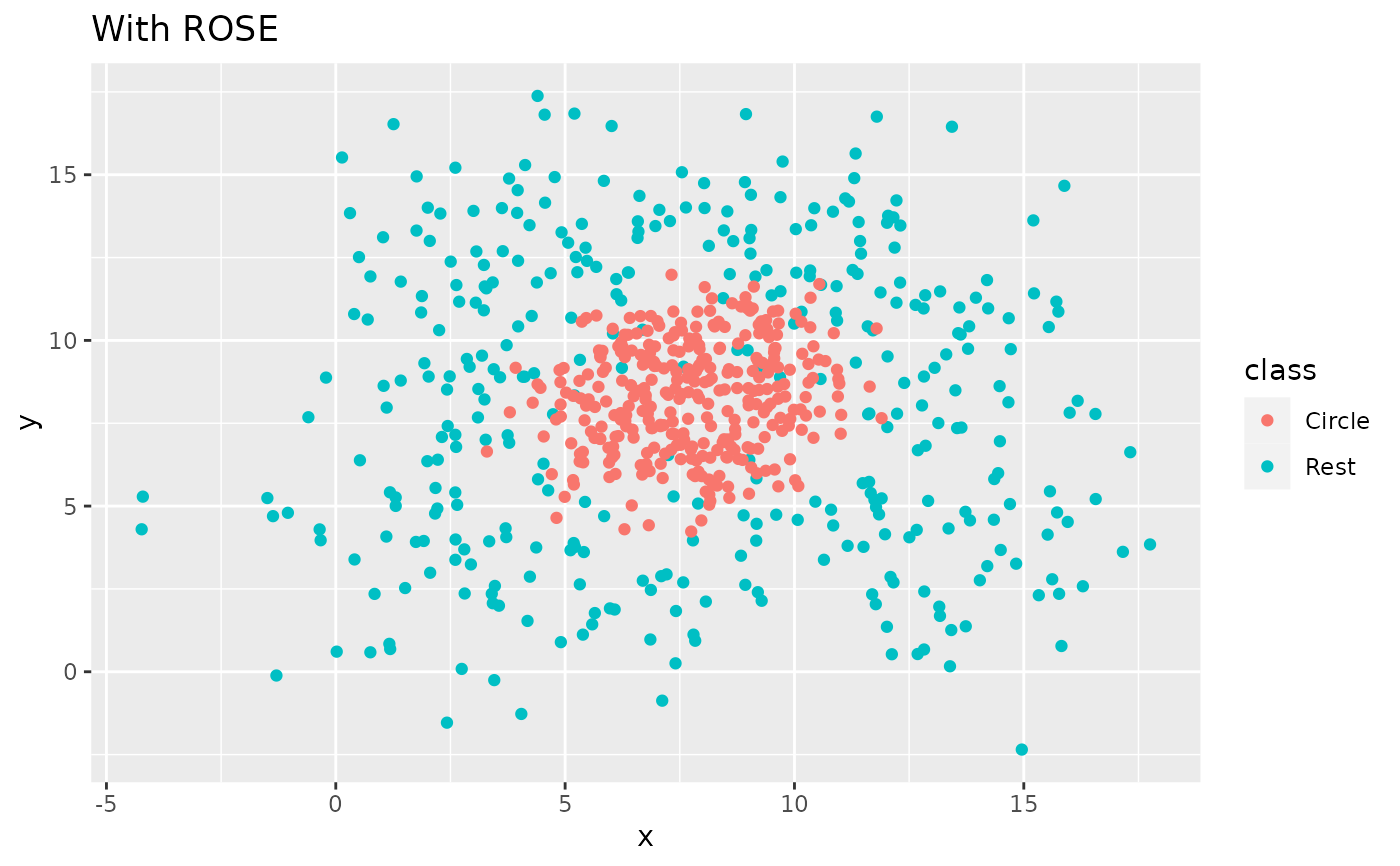
 recipe(class ~ x + y, data = circle_example) |>
step_rose(class) |>
prep() |>
bake(new_data = NULL) |>
ggplot(aes(x, y, color = class)) +
geom_point() +
labs(title = "With ROSE")
recipe(class ~ x + y, data = circle_example) |>
step_rose(class) |>
prep() |>
bake(new_data = NULL) |>
ggplot(aes(x, y, color = class)) +
geom_point() +
labs(title = "With ROSE")